A visualization method for displaying the structure of process bigraphs
Project description
Bigraph-viz
Bigraph-viz is an easy-to-use plotting tool for compositional bigraph schema, built on top of Graphviz.
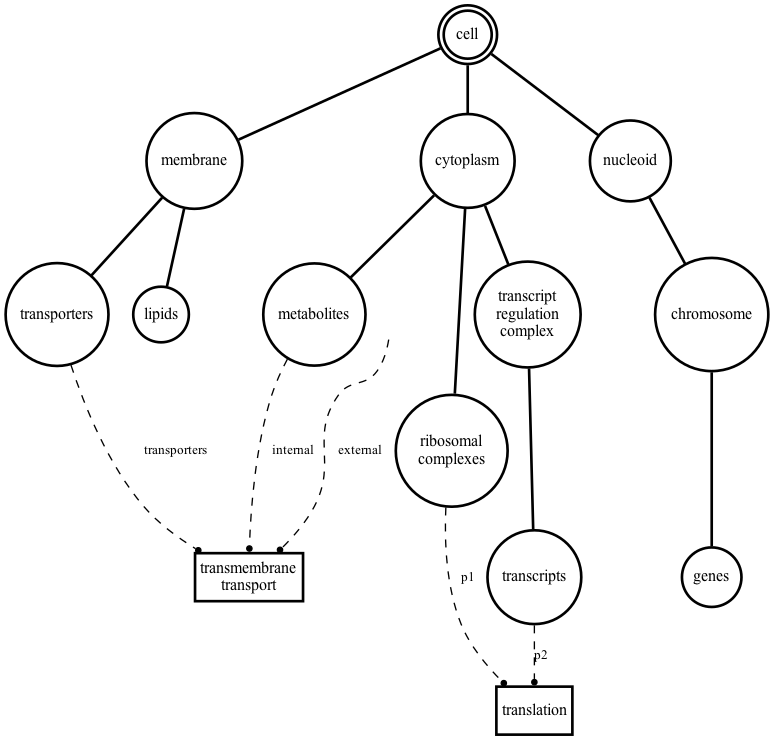
Compositional Schemas and Milner Bigraphs
Compositional Bigraph Schemas (CBS) are based on a mathematical formalism introduced by Robin Milner in 2009 , and used in Vivarium. Bigraphs consist of networks with embeddable nodes that can be placed within other nodes and dynamically restructured. CBS reimagines the bigraph concept. Variables are contained within Stores (the circles in the figure), which can be embedded in a hierarchy, as shown by the dark edges. Instead of Milner's hyperedges, CBS employs Processes (the rectangles) which connect via wires (dashed edges) to shared variables within the Stores. Processes and Stores form a type of bipartite graph, as illustrated by the dashed edges.
In Vivarium, the CBS is used to structure a modular multiscale simulation. Bigraph-viz is part of an effort to standardize this structure, so that it can be used as an exchange format for multiscale simulations.
Getting Started
Dependencies
Before installing bigraph-viz, make sure you have Graphviz.
Ubuntu/Debian
sudo apt-get install -y graphviz
macOS
brew install graphviz
Windows
Download and install the Graphviz installer from the official website. Make sure to add the Graphviz bin directory to your system's PATH.
Installation
Once Graphviz is installed, you can install bigraph-viz using pip:
pip install bigraph-viz
Notebooks
All notebooks are tested in CI and published as HTML to GitHub Pages: Browse all notebooks
| Notebook | Description |
|---|---|
| Basics | Tutorial covering stores, processes, wires, and composites |
| Basics2 | Extended basics with cell structure example |
| Basics Pres | Presentation-style examples |
| Bigraph Example | Transcription/translation and multicellular models |
| Cell Atlas | Kidney glomerulus model |
| Ecoli | Whole-cell E. coli wiring diagram |
| Ecoli hybrid | E. coli hybrid framework |
| Format | Styling options for bigraph figures |
| Gut Microbiome | Multi-region gut model with cdFBA-CRM |
| Ccb | CCB model (source only) |
License
Bigraph-viz is open-source software released under the Apache 2 License.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file bigraph_viz-2.0.3.tar.gz.
File metadata
- Download URL: bigraph_viz-2.0.3.tar.gz
- Upload date:
- Size: 37.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
82f7a50a5d135c07fa4ab28a6f768575d0bba9f09b9fe870996b4b53b9c87274
|
|
| MD5 |
a70f21d58f3bbaa7c9d5a5471ea8b91e
|
|
| BLAKE2b-256 |
aee3aae87e36b5424fcf5b210961ad6a09a89defddf8d7d01552d51ff57376eb
|
Provenance
The following attestation bundles were made for bigraph_viz-2.0.3.tar.gz:
Publisher:
release.yml on vivarium-collective/bigraph-viz
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
bigraph_viz-2.0.3.tar.gz -
Subject digest:
82f7a50a5d135c07fa4ab28a6f768575d0bba9f09b9fe870996b4b53b9c87274 - Sigstore transparency entry: 1508668940
- Sigstore integration time:
-
Permalink:
vivarium-collective/bigraph-viz@3b9115b9166748ae41bc0a2e7b38f3b1d6f29009 -
Branch / Tag:
refs/tags/v2.0.3 - Owner: https://github.com/vivarium-collective
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@3b9115b9166748ae41bc0a2e7b38f3b1d6f29009 -
Trigger Event:
push
-
Statement type:
File details
Details for the file bigraph_viz-2.0.3-py3-none-any.whl.
File metadata
- Download URL: bigraph_viz-2.0.3-py3-none-any.whl
- Upload date:
- Size: 36.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c464309f861471cdf3661a0b4078566137f4b5be370fa9fecd4848454a8c7884
|
|
| MD5 |
7f3a623888f05b79387c0ebb4f9ee274
|
|
| BLAKE2b-256 |
cc5a9b55539d08a46418b2389e18fe01229584e0f0b1139d10c783e9491ace9c
|
Provenance
The following attestation bundles were made for bigraph_viz-2.0.3-py3-none-any.whl:
Publisher:
release.yml on vivarium-collective/bigraph-viz
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
bigraph_viz-2.0.3-py3-none-any.whl -
Subject digest:
c464309f861471cdf3661a0b4078566137f4b5be370fa9fecd4848454a8c7884 - Sigstore transparency entry: 1508669020
- Sigstore integration time:
-
Permalink:
vivarium-collective/bigraph-viz@3b9115b9166748ae41bc0a2e7b38f3b1d6f29009 -
Branch / Tag:
refs/tags/v2.0.3 - Owner: https://github.com/vivarium-collective
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@3b9115b9166748ae41bc0a2e7b38f3b1d6f29009 -
Trigger Event:
push
-
Statement type:














