Custom time-series export of LAIe/FAPAR/FCOVER with YAML params.
Project description
gee-biophys
Custom temporal exports of biophysical variable retrievals (LAIe, FAPAR, FCOVER) from Sentinel-2 imagery.
This tool enables users to define custom time windows, temporal frequency, regions, and export settings for Sentinel-2–based biophysical variable retrievals using a simple YAML configuration file. It provides both:
- a Python API, and
- a Command-Line Interface (CLI):
gee-biophys.
Overview
gee-biophys enables flexible and reproducible export of time-series vegetation biophysical maps, addressing the need for user-defined temporal aggregation of Sentinel-2–derived biophysical variable retrievals.
The tool builds on the PROSAIL-based s2biophys retrieval framework, producing consistent global estimates of:
- Effective Leaf Area Index (LAIe)
- Fraction of Absorbed Photosynthetically Active Radiation (FAPAR)
- Fractional Vegetation Cover (FCOVER)
Each export includes:
mean— aggregated trait valuestdDev— within-period variabilitycount— number of valid observations
Features
- Flexible temporal definitions: fixed (e.g. monthly, quarterly) or custom seasonal windows
- Spatial inputs via bounding box or GeoJSON geometry
- Modular YAML configuration for reproducibility
- Exports directly to Google Earth Engine assets, Google Drive, or Google Cloud Storage
- Available as both a Python package and CLI tool
- Supports two models: s2biophys and SL2P models
Installation
Using pip
Create a clean virtual environment (e.g. using conda or venv). Recommended Python version 3.12:
conda create -n [ENV_NAME] python=3.12
conda activate [ENV_NAME]
pip install gee-biophys
Alternative installation — Install from source
conda create -n [ENV_NAME] python=3.12
conda activate [ENV_NAME]
git clone https://github.com/speckerf/gee_biophys.git
cd gee_biophys
pip install -e .
Usage
Start Exports
1. Command-line interface
gee-biophys --config path_to_yaml.yaml (--public)
- The
--config(or-c) argument is required and must point to a valid YAML configuration file. - The
--publicargument is optional and can be specified if the createImageCollectionshould be public to all GEE-users.
2. Python interface
from gee_biophys.cli import run_pipeline
run_pipeline(config='path_to_yaml.yaml')
Wait for exports to finish:
The tool will start an Earth Engine export task for each exported time period. So the temporal frequency and the total time window determine the number of export tasks to execute.
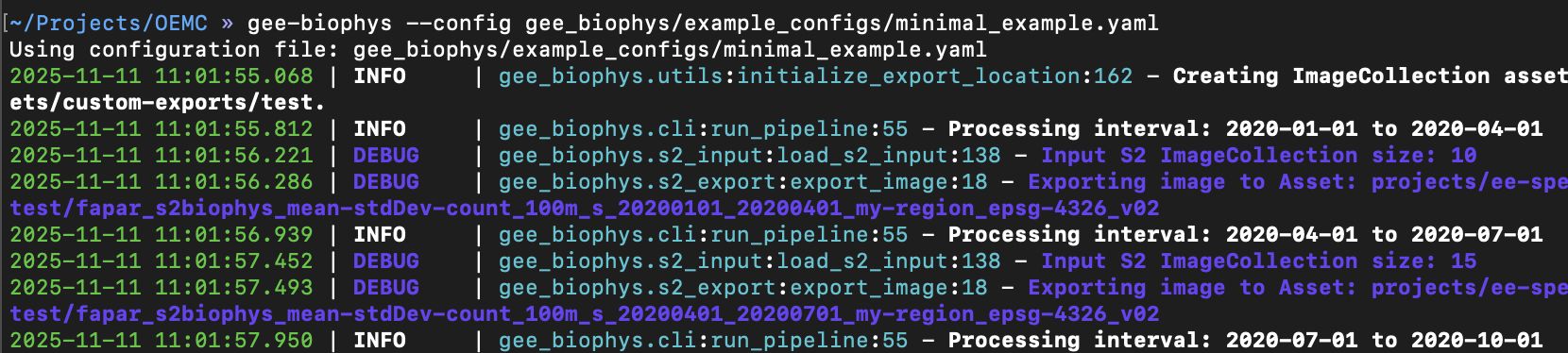
Check the progress of the exports in the code editor directly:
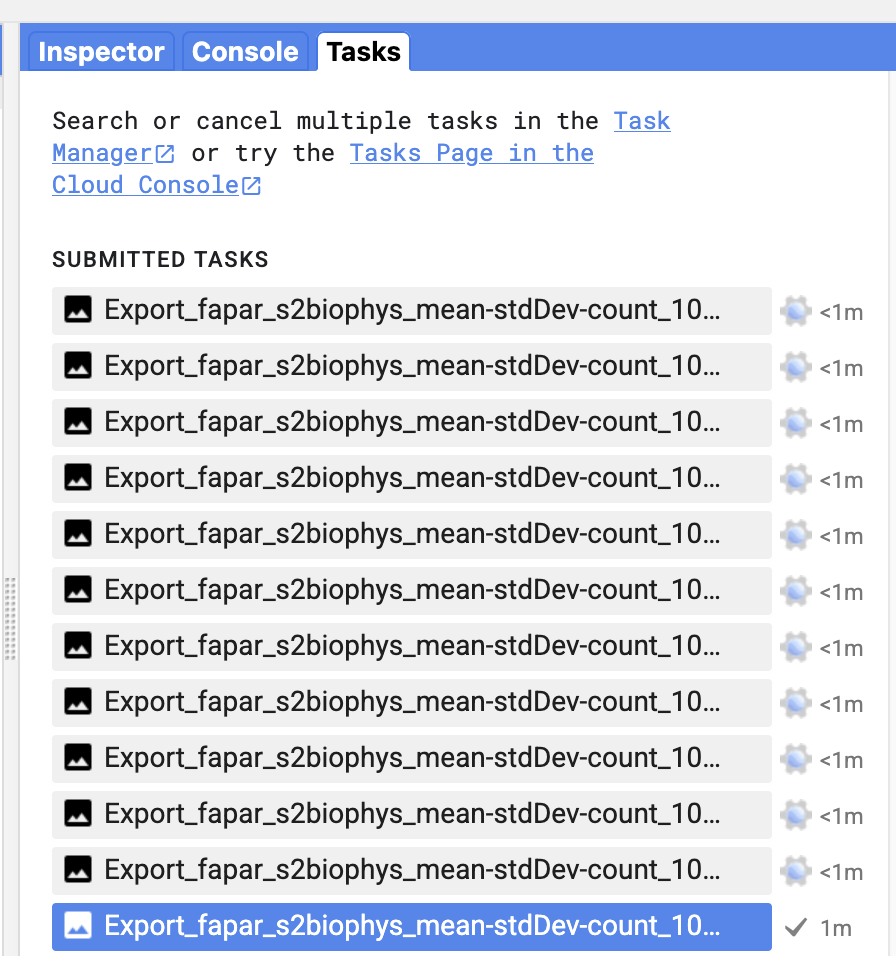
Alternatively, check progress directly using the logged task ID: earthengine task info [TASK_ID]
Visualize results
To visualize results, please use the following earth-engine app: here. Note that this requires that the option --public was set when running gee-biophys.
Alternatively, the source code of the app can also be directly to visualize a non-public ImageCollection.
Example Configuration
Example configuration files are provided in https://github.com/speckerf/gee_biophys/tree/main/example_configs.
Minimal Example Config
# ============ GEE-Biophys ================
# Minimal Example (small exports for testing)
# ==============================================
spatial:
type: bbox
bbox: [7.1, 46.1, 7.2, 46.2]
region_name: my-region
temporal:
start: "2020-01-01"
end: "2023-01-01"
cadence:
type: fixed
interval: quarterly
variables:
model: s2biophys
variable: fapar
export:
destination: asset
collection_path: "projects/ee-speckerfelix/assets/custom-exports/test" # CHANGE
project_id: "ee-speckerfelix" # CHANGE
crs: "EPSG:4326" # EPSG code or LOCAL_UTM (automatic)
scale: 100
max_pixels: 100_000_000_000
options:
max_cloud_cover: 50
csplus_band: cs
cs_plus_threshold: 0.70
clip_min_max: true # if true, clips predictions to (0, 1) for fapar and fcover, and (0, 8) for laie/lai
version: "v02"
Reference Template
A fully commented reference configuration file is available here. A minimal example can be found here. Please check some example configuration files available here.
Configuration Structure
| Section | Purpose | Example |
|---|---|---|
spatial |
Defines the area of interest (bbox or GeoJSON) | [minLon, minLat, maxLon, maxLat] |
temporal |
Sets start/end dates and cadence (fixed or seasonal) | quarterly, monthly, yearly |
variables |
Selects the biophysical variable and retrieval model | fapar, laie, fcover |
export |
Configures export target and GEE project, CRS, spatial resolution, etc. | e.g. projects/ee-user/assets/... |
options |
Controls cloud masking and thresholds | max cloud cover, CS+ threshold |
version |
Records model version for reproducibility | "v02" |
Note: If you want to find a good balance between strict cloud masking and having enough cloud-free pixels for analysis, you can use the following script to explore and tune the three key parameters—CloudScore+ band, CloudScore+ threshold, and maximum CLOUDY_PIXEL_PERCENTAGE from the Sentinel-2 metadata: https://code.earthengine.google.com/716bee247685008f34b49c63d32b8447
File Naming Convention
Each exported asset or file follows a standardized and descriptive naming scheme generated automatically from the configuration parameters. The convention ensures reproducibility, traceability, and easy identification of spatial, temporal, and model settings.
The filename (or system:index in GEE) is constructed as:
{variable}_{model}_{bands}_{scale}m_s_{start}_{end}_{region-name}_{crs}_{version}
Example
fapar_s2biophys_mean-stdDev-count_100m_s_20200101_20201231_my-region_epsg-4326_v02
Note that exports with the same system:index will fail when writing to Earth Engine assets, as each asset ID must be unique, whereas they will overwrite existing files when exporting to Google Drive or Google Cloud Storage.
Citation
If you use gee-biophys in your research, please cite the associated publication (forthcoming):
Felix Specker, Anna K. Schweiger, Jean-Baptiste Féret et al. Advancing Ecosystem Monitoring with Global High-Resolution Maps of Vegetation Biophysical Properties, 01 April 2025, PREPRINT (Version 1) available at Research Square [https://doi.org/10.21203/rs.3.rs-6343364/v1]
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file gee_biophys-1.3.0.tar.gz.
File metadata
- Download URL: gee_biophys-1.3.0.tar.gz
- Upload date:
- Size: 42.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.9.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
869ef0fcc68e46b12e5aacd98c3a10f6c2710d518d49e9395e988bab11225a3b
|
|
| MD5 |
71e1a5ff6e162a7e5aa1fdc65dd2ea7f
|
|
| BLAKE2b-256 |
0953956f6f0e63fef6ad6066b605c87da7da12402524334fbf08874b2be87bf0
|
File details
Details for the file gee_biophys-1.3.0-py3-none-any.whl.
File metadata
- Download URL: gee_biophys-1.3.0-py3-none-any.whl
- Upload date:
- Size: 42.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.9.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d37e0c831792ebbeea31d6b6e82acdad2c62889e678c77b68fa3c7783c21bc56
|
|
| MD5 |
19f43003a0233f4ace8da78b47f8d0e1
|
|
| BLAKE2b-256 |
e458e9e58a3d139456b514abb5fe4b407df02d81007c73f7423b41921a525374
|











